To address this question, we mainly use Arabidopsis thaliana as a model system and combine molecular biology, genetic, epigenomic, biochemistry, imaging and bioinformatics approaches.
Networks: Our research is integrated into local initiatives such as the CAP20-25 project International research Centre 1 (CIR1) on Sustainable Agro-ecosystems and CIR3 on mobility, national networks such as the GDRs (Groupements de Recherche) EPIPLANT, the ADN&G, the IMABIO, or the RtmFM network, European projects such as Marie Curie Doctoral Networks, and international networks through our involvement in the Society of Experimental Biology (SEB).
Teaching and Outreach: We contribute actively to the life of Clermont Auvergne University through our implication in the Unité de Formation et de Recherche (UFR) Biology and our contribution to courses in genetics in the Bachelor of Life Sciences, epigenetics in the Master of Health Biology and in the Master of Plant Biology, and bioinformatics in the Master of Bioinformatics. We are also engaged in the dissemination of scientific culture through our participation in outreach events through the local association astu’science.
Our research is supported by EU MSCA DN EpiSeedLink and AGILE, 4D-Heat [ANR-21-CE20-0036], SeedChrom [ANR-22-CE20-0028], EpiLinks [ANR-22-CE20-0001], NUCLINV [ANR-24-CE20-5404] and His2AVar and through funding from the CAP20-25 CIR1 and CIR3.
Funders

Research
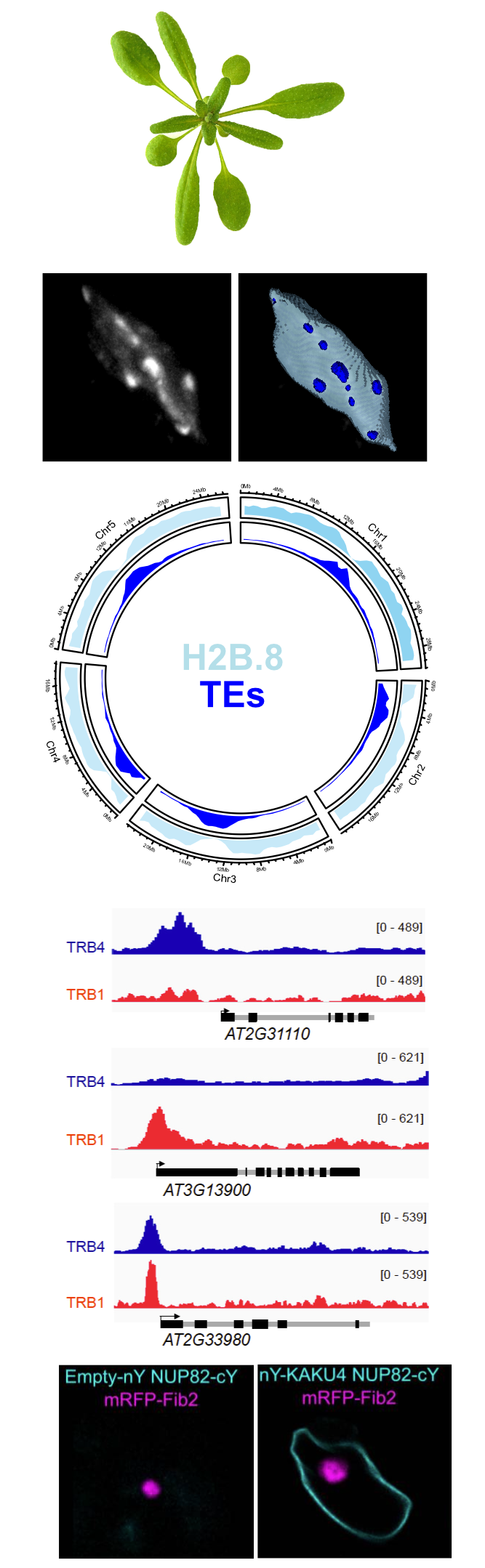
We currently focus on three research areas:
Histone variants and chaperones
Precise dynamics of histone proteins is critical for all DNA-template based processes that take place in the context of chromatin. A network of histone chaperone proteins therefore tightly coordinates the incorporation of histone variants into the nucleosomes. We study how the incorporation of different histone variants changes nucleosome properties, influences higher-order chromatin organization and affects gene activity. We focus on the transition from seed to seedling, a key developmental transition in plants that involves major changes in gene expression and chromatin organization.
A. Probst, S. Cotterell, S. Le Goff, L. Simon, R. Kaur, S. Paltrinieri, G. Sireta. M. Verdier
Coordinating Polycomb activity through GH1 domain proteins
Polycomb group (PcG) proteins play a central role in controlling developmental genes that govern growth, differentiation, and morphogenesis in eukaryotes. Proteins with a globular histone H1 (GH1) domain contribute to H3K27me3 homeostasis either by favoring the establishment of a repressive chromatin structure or through direct recruitment of the Polycomb Repressive 2 (PRC2) complexes. We aim to understand how Telomere Repeat Binding (TRB) proteins, GH1-High mobility group proteins (GH1-HMGA) and the linker histone H1 work together to fine-tune Polycomb activity and to regulate gene expression.
S. Amiard, E. Vanrobays, S. Tutois, L. Feit, H. El Idrissi Moubtassim
Assessing variability in chromatin organization at single cell level
Precise and quantitative evaluation of chromatin and nuclear organization at the level of a single cell single is a prerequisite for uncovering structural and functional principles at unprecedented spatial and temporal scale. To assess chromatin organization at the single-cell level, we develop advanced imaging methods combined with deep-learning image analysis tools for 3D image segmentation and cell tracking in different model systems.
C. Tatout, C. Desset, S. Safarbati
Former Team Members
Shimlal Ayilalath, PhD, 2022-25
Aurelia Stevens, Assistant Professor, 2018-24
Guillaume Mougeot, PhD, 2021-23
Sarah Mermet, PhD, 2019-22
Tristan Dubos, PhD, 2018-21
Gwénaëlle Détourné, PhD, 2015-19
Maxime Voisin, PhD, 2013-17
Céline Duc, Postdoc, 2010-14
Lauriane Simon, PhD, 2012-16
Axel Poulet, PhD, 2012-16
Mélanie Thomas, PhD, 2011-14
Matthias Benoit, PhD, 2010-14
Members
Publications
New three-dimensional nuclear morphometry tool quantifies impact of slow freezing on sperm hypercondensed chromatin.
Published on 01 Mar 2026 in Andrology , vol. 14 - pp 809-821
Carcy O, Desset S , Dubos T , Puceat C, Andrieux M, Compagnon M, Sireta G , Pereira B, Cachin F, Probst AV , Tatout C , Brugnon F, Pons-Rejraji H
Sorting F. graminearum core effector candidates shows multiple fungal proteins that target the wheat cell nucleus during Fusarium Head Blight
Published on 09 Feb 2026 in Journal of Experimental Botany , vol. erag067
Ayilalath S. , Faurie L., Vanrobays E. , Rocher F., Loizeau L., Philippe G., Javelle M., Bosio M., Sallaud C., Tatout C. , Bonhomme L.
CRESCENT, a comprehensive RNA-Seq expression, splicing, and coding/non-coding element network tool.
Published on 21 Jan 2026 in BMC bioinformatics , vol. 27 - pp 43
Sireta G , Cueff G, Darbot V, Lefebvre M, Amiard S , Probst AV , Tatout C
Enhanced lysosomal glycogen breakdown is associated with liver tumorigenesis in glycogen storage disease type III
Published on 01 Dec 2025 in JHEP Reports , vol. 8 - pp 1-13
Valle Montalvo-Romeral, Louisa Jauze, Gwendoline Perrot, Mouna Amaouche, Antoine Gardin, Araceli Aguilar González, Alicia Leblond , Carine Zitoun-Ardon , Félicie Evrard , Jérémie Cosette , Christophe Tatout , Fanny Bordier , Emilie Bertil-Froidevaux, Christophe Georger , Laetitia van Wittenberghe , Valérie Paradis , Simon Gay , Fanny Dujardin , François Maillot, Amandine Gautier-Stein , Gilles Mithieux, Edoardo Malfatti, Charlotte Mussini , Philippe Labrune , Alicia Mayeuf-Louchart, Giuseppe Ronzitti, Fabienne Rajas
The mid-SUN-POD1 complex ensures the structural integrity of ER bodies required for herbivore defense in Arabidopsis.
Published on 01 Oct 2025 in The New phytologist
Ikeda F, Ohtsubo T, Nakagawa S, Suzuki T, Miyoshi N, Vanrobays E , Tatout C , Shimada T, Tamura K
Hibernating brown bear serum modulates the balance of TGF-β and BMP pathways in human muscle cells.
Published on 19 Sep 2025 in The international journal of biochemistry & cell biology - pp 106864
Richard C, Pourpe C, Fourneaux G, Cueff G, Parry L, Coudy-Gandilhon C, Kindberg J, Evans AL, Miller A, Gauquelin-Koch G, Tatout C , Polge C, Taillandier D, Bertile F, Lefai E, Combaret L
Replication-independent eviction of H2B.8 reveals chromatin reprogramming during seed imbibition
Published on 23 Aug 2025 in bioRxiv
Simon L , Paltrinieri S , Verdier M , Wang Q, Cotterell S , Latrasse D, Maugarny-Cales A, Sireta G , Desset S , Amiard S , Bailly C, Tamura K, Benhamed M, Tatout C , Le Goff S , Probst AV
Ubiquitin-dependent proteolysis of KNL2 driven by APC/CCDC20 is critical for centromere integrity and mitotic fidelity
Published on 25 Jun 2025 in The Plant Cell , vol. koaf164
Kalidass M Jarubula VG, Ratnikava M, Chandra JR, Le Goff S , Probst AV , Esposito S, Grasser KD, Bruckmann A, Gagneux JF, Prosée RF, Rutten T, Schubert V, Demidov D, Lechner E, Steiner FA, Genschik P, Lermontova I
Plant epigenetics: controlling genome expression to integrate developmental and environmental cues
Published on 17 Jun 2025 in J Exp Bot , vol. 76 - pp 2389-2393
Catoni M, Lechon Gomez T, Probst AV
Sorting F. graminearum core effector candidates shows multiple fungal proteins that target the wheat cell nucleus during Fusarium Head Blight
Published on 08 Feb 2025 in bioRxiv
Ayilalath S , Faurie L , Vanrobays E , Rocher F, Loizeau L, Philippe G, Javel M, Bosio M, Sallaud C, Tatout C , Bonhomme L
Characterizations of Nuclear Organization and Chromatin Modifications in Mature Embryos from Arabidopsis Dry Seeds.
Published on 01 Jan 2025 in Methods in molecular biology (Clifton, N.J.) , vol. 2873 - pp 247-262
The dual trxG/PcG protein ULTRAPETALA1 modulates H3K27me3 and directly enhances POLYCOMB REPRESSIVE COMPLEX 2 activity for fine-tuned reproductive transitions
Published on 24 Oct 2024 in bioRxiv
Geshkovski V, Engelhorn J, Izquierdo JB, Laroussi H, Thouly C, Turchi L, Le Masson M, Thévenon E, Petitalot A, Simon L , Desset S , Michl-Holzinger P, Parrinello H, Grasser KD, Probst AV , Margueron R, Vachon G, Kadlec J, Carles CC
The histone chaperones ASF1 and HIRA are required for telomere length and 45S rDNA copy number homeostasis
Published on 14 Oct 2024 in The Plant Journal , vol. 120 - pp 1125-1141
Machelová A, Dadejová MN, Franek M, Mougeot G , Simon L , Le Goff S , Duc C , Bassler J, Demko M, Schwarzerová J, Desset S , Probst AV , Dvořáčková M
Biom3d, a modular framework to host and develop 3D segmentation methods
Published on 25 Jul 2024 in bioRxiv
Mougeot G , Safarbati S , Alégot H , Pouchin P , Field N, Almagro S, Pery E, Probst AV , Tatout C , Evans DE, Graumann K, Chausse F, Desset S
Maintenance and dynamic reprogramming of chromatin organization during development.
Published on 01 May 2024 in The Plant journal , vol. 118 - pp 657-670
The TELOMERE REPEAT BINDING proteins TRB4 and TRB5 function as transcriptional activators of PRC2-controlled genes to regulate plant development.
Published on 01 Apr 2024 in Plant communications - pp 100890
Amiard S , Feit L , Vanrobays E , Simon L , Le Goff S , Loizeau L , Wolff L, Butter F, Bourbousse C, Barneche F, Tatout C , Probst AV
Evolutionarily conserved protein motifs drive interactions between the plant nucleoskeleton and nuclear pores
Published on 21 Sep 2023 in The Plant Cell
Mermet S , Voisin M , Mordier J , Dubos T , Tutois S , Tuffery P, Baroux C, Tamura K, Probst AV , Vanrobays E , Tatout C
Histone H1 protects telomeric repeats from H3K27me3 invasion in Arabidopsis
Published on 28 Jul 2023 in Cell Reports , vol. 42 - pp 112894
Teano G, Concia L, Wolff L, Carron L, Biocanin I, Adamusová K, Miloslava Fojtová, Michael Bourge, Amira Kramdi, Vincent Colot, Ueli Grossniklaus, Bowler C, Baroux C, Carbone A, Probst AV , Procházková Schrumpfová P, Fajkus J, Amiard S , Grob S, Bourbousse C, Barneche F
HSFA1a modulates plant heat stress responses and alters the 3D chromatin organization of enhancer-promoter interactions
Published on 28 Jan 2023 in Nat. Comm. , vol. 14 - pp 469
Huang Y, An J, Sircar S, Bergis C, Lopes CD, He X, Da Costa B, Tan FQ, Bazin J, Antunez-Sanchez J, Mammarella MF, Devani RS, Brik-Chaouche R, Bendahmane A, Frugier F, Xia C, Rothan C, Probst AV , Mohamed Z, Bergounioux C, Delarue M, Zhang Y, Zheng S, Crespi M, Fragkostefanakis S, Mahfouz MM, Ariel F, Gutierrez-Marcos J, Raynaud C, Latrasse D, Benhamed M
Image analysis workflows to reveal the spatial organization of cell nuclei and chromosomes.
Published on 30 Dec 2022 in Nucleus (Austin, Tex.) , vol. 13 - pp 277-299
Randall RS, Jourdain C, Nowicka A, Kaduchová K, Kubová M, Ayoub MA, Schubert V, Tatout C , Colas I, Kalyanikrishna, Desset S , Mermet S , Boulaflous-Stevens A, Kubalová I, Mandáková T, Heckmann S, Lysak MA, Panatta M, Santoro R, Schubert D, Pecinka A, Routh D, Baroux C
Histone H1 protects telomeric repeats from H3K27me3 invasion in Arabidopsis
Published on 06 Dec 2022 in bioRxiv
Teano G, Concia L, Wolff L, Carron L, Biocanin I, Adamusová K, Fojtová M, Bourge M, Kramdi A, Colot V, Grossniklaus U, Bowler C, Baroux C, Carbone A, Probst AV , Procházková Schrumpfová P, Fajkus J, Amiard S , Grob S, Bourbousse C, Barneche F
Easing batch image processing from OMERO: a new toolbox for ImageJ.
Published on 24 Nov 2022 in F1000Research , vol. 11 - pp 392
Pouchin P , Zoghlami R, Valarcher R , Delannoy M, Carvalho M, Belle C, Mongy M, Desset S , Brau F
Deposition and eviction of histone variants define functional chromatin states in plants.
Published on 30 Oct 2022 in Current opinion in plant biology , vol. 69 - pp 102266
NODeJ: an ImageJ plugin for 3D segmentation of nuclear objects.
Published on 06 Jun 2022 in BMC bioinformatics , vol. 23 - pp 216
Dubos T , Poulet A , Thomson G, Péry E, Chausse F, Tatout C , Desset S , van Wolfswinkel JC, Jacob Y
Deep learning — promises for 3D nuclear imaging: a guide for biologists.
Published on 01 Apr 2022 in Journal of cell science , vol. 135
Mougeot G , Dubos T , Chausse F, Péry E, Graumann K, Tatout C , Evans DE, Desset S
ANCHOR: A Technical Approach to Monitor Single-Copy Locus Localization in Planta.
Published on 06 Jul 2021 in Frontiers in plant science , vol. 12 - pp 677849
Meschichi A, Ingouff M, Picart C, Mirouze M, Desset S , Gallardo F, Bystricky K, Picault N, Rosa S, Pontvianne F
Polycomb-dependent differential chromatin compartmentalization determines gene coregulation in Arabidopsis.
Published on 03 Jun 2021 in Genome research , vol. 31 - pp 1230-44
Huang Y, Sicar S, Ramirez-Prado JS, Manza-Mianza D, Antunez-Sanchez J, Brik-Chaouche R, Rodriguez-Granados NY, An J, Bergounioux C, Mahfouz MM, Hirt H, Crespi M, Concia L, Barneche F, Amiard S , Probst AV , Gutierrez-Marcos J, Ariel F, Raynaud C, Latrasse D, Benhamed M
Gene dosage compensation of rRNA transcript levels in Arabidopsis thaliana lines with reduced ribosomal gene copy number
Published on 31 May 2021 in The Plant Cell , vol. 33 - pp 1135-1150
Lopez FB, Fort A, Tadini L, Probst AV , McHale M, Friel J, Ryder P, Pontvianne FDR, Pesaresi P, Sulpice R, McKeown P, Brychkova G, Spillane C
The Histone Chaperone HIRA Is a Positive Regulator of Seed Germination.
Published on 14 Apr 2021 in International journal of molecular sciences , vol. 22
Layat E , Bourcy M, Cotterell S , Zdzieszyńska J, Desset S , Duc C , Tatout C , Bailly C, Probst AV
Automated 3D bio-imaging analysis of nuclear organization by NucleusJ 2.0.
Published on 30 Dec 2020 in Nucleus (Austin, Tex.) , vol. 11 - pp 315-329
Dubos T , Poulet A , Gonthier-Gueret C, Mougeot G , Vanrobays E , Li Y , Tutois S , Pery E, Chausse F, Probst AV , Tatout C , Desset S
Untangling chromatin interactions.
Published on 17 Aug 2020 in Journal of experimental botany , vol. 71 - pp 5115-5118
Similar yet critically different: the distribution, dynamics and function of histone variants.
Published on 17 Aug 2020 in Journal of experimental botany , vol. 71 - pp 5191-5204
Probst AV , Desvoyes B, Gutierrez C
The H3 histone chaperone NASP(SIM3) escorts CenH3 in Arabidopsis.
Published on 30 Jan 2020 in The Plant Journal , vol. 101 - pp 71-86
Le Goff S , Keçeli BN, Jeřábková H, Heckmann S, Rutten T, Cotterell S , Schubert V, Roitinger E, Mechtler K, Franklin FCH, Tatout C , Houben A, Geelen D, Probst AV , Lermontova I
Probing the 3D architecture of the plant nucleus with microscopy approaches: challenges and solutions.
Published on 30 Dec 2019 in Nucleus (Austin, Tex.) , vol. 10 - pp 181-212
Dumur T, Duncan S, Graumann K, Desset S , Randall RS, Scheid OM, Prodanov D, Tatout C , Baroux C
Looking At the Past and Heading to the Future: Meeting Summary of the 6(th) European Workshop on Plant Chromatin 2019 in Cologne, Germany.
Published on 07 Feb 2019 in Frontiers in plant science , vol. 10 - pp 1795
Moreno-Romero J, Probst AV , Trindade I, Kalyanikrishna, Engelhorn J, Farrona S
Replication-coupled histone H3.1 deposition determines nucleosome composition and heterochromatin dynamics during Arabidopsis seedling development.
Published on 30 Jan 2019 in The New phytologist , vol. 221 - pp 385-398
Benoit M , Simon L , Desset S , Duc C , Cotterell S , Poulet A , Le Goff S , Tatout C , Probst AV
Spatial organization of chromosome territories in the interphase nucleus of trisomy 21 cells.
Published on 30 Jun 2018 in Chromosoma , vol. 127 - pp 247-259
Kemeny S, Tatout C , Salaun G, Pebrel-Richard C, Goumy C, Ollier N, Maurin E, Pereira B, Vago P, Gouas L
Meeting report – INDEPTH kick-off meeting.
Published on 25 Jun 2018 in Journal of cell science , vol. 131
The Linker Histone GH1-HMGA1 Is Involved in Telomere Stability and DNA Damage Repair.
Published on 30 May 2018 in Plant physiology , vol. 177 - pp 311-327
Charbonnel C, Rymarenko O, Da Ines O , Benyahya F, White CI, Butter F, Amiard S
Genetic and epigenetic variation in 5S ribosomal RNA genes reveals genome dynamics in Arabidopsis thaliana.
Published on 06 Apr 2018 in Nucleic acids research , vol. 46 - pp 3019-3033
Simon L , Rabanal FA, Dubos T , Oliver C, Lauber D , Poulet A , Vogt A, Mandlbauer A, Le Goff S , Sommer A, Duborjal H, Tatout C , Probst AV
A Compendium of Methods to Analyze the Spatial Organization of Plant Chromatin.
Published on 01 Jan 2018 in Methods in molecular biology (Clifton, N.J.) , vol. 1675 - pp 397-418
High-Affinity LNA-DNA Mixmer Probes for Detection of Chromosome-Specific Polymorphisms of 5S rDNA Repeats in Arabidopsis thaliana.
Published on 01 Jan 2018 in Methods in molecular biology (Clifton, N.J.) , vol. 1675 - pp 481-491
Quantitative 3D Analysis of Nuclear Morphology and Heterochromatin Organization from Whole-Mount Plant Tissue Using NucleusJ.
Published on 01 Jan 2018 in Methods in molecular biology (Clifton, N.J.) , vol. 1675 - pp 615-632
Investigation of Nuclear Periphery Protein Interactions in Plants Using the Membrane Yeast Two-Hybrid (MbY2H) System.
Published on 01 Jan 2018 in Methods in molecular biology (Clifton, N.J.) , vol. 1840 - pp 221-235
Computational Methods for Studying the Plant Nucleus.
Published on 01 Jan 2018 in Methods in molecular biology (Clifton, N.J.) , vol. 1840 - pp 205-219
Poulet A , Zhou X, Tamura K, Meier I, Tatout C , Graumann K, Evans DE
Arabidopsis ATRX Modulates H3.3 Occupancy and Fine-Tunes Gene Expression.
Published on 30 Jul 2017 in The Plant cell , vol. 29 - pp 1773-1793
Duc C , Benoit M , Détourné G , Simon L , Poulet A , Jung M, Veluchamy A, Latrasse D, Le Goff S , Cotterell S , Tatout C , Benhamed M, Probst AV
The LINC complex contributes to heterochromatin organisation and transcriptional gene silencing in plants.
Published on 01 Feb 2017 in Journal of cell science , vol. 130 - pp 590-601
Poulet A , Duc C , Voisin M , Desset S , Tutois S , Vanrobays E , Benoit M , Evans DE, Probst AV , Tatout C
Exploring the evolution of the proteins of the plant nuclear envelope.
Published on 02 Jan 2017 in Nucleus (Austin, Tex.) , vol. 8 - pp 46-59
A novel family of plant nuclear envelope-associated proteins.
Published on 30 Oct 2016 in Journal of experimental botany , vol. 67 - pp 5699-5710
Pawar V, Poulet A , Détourné G , Tatout C , Vanrobays E , Evans DE, Graumann K
Structure and Function of Centromeric and Pericentromeric Heterochromatin in Arabidopsis thaliana.
Published on 30 Nov 2015 in Frontiers in plant science , vol. 6 - pp 1049
Stress-induced structural changes in plant chromatin.
Published on 30 Oct 2015 in Current opinion in plant biology , vol. 27 - pp 8-16
Probst AV , Mittelsten Scheid O
NucleusJ: an ImageJ plugin for quantifying 3D images of interphase nuclei.
Published on 01 Apr 2015 in Bioinformatics (Oxford, England) , vol. 31 - pp 1144-6
Poulet A , Arganda-Carreras I, Legland D, Probst AV , Andrey P, Tatout C
The histone chaperone complex HIR maintains nucleosome occupancy and counterbalances impaired histone deposition in CAF-1 complex mutants.
Published on 30 Mar 2015 in The Plant journal , vol. 81 - pp 707-22
Duc C , Benoit M , Le Goff S , Simon L , Poulet A , Cotterell S , Tatout C , Probst AV
Marker gene tethering by nucleoporins affects gene expression in plants.
Published on 01 Jan 2015 in Nucleus (Austin, Tex.) , vol. 6 - pp 471-8
Smith S, Galinha C, Desset S , Tolmie F, Evans D, Tatout C , Graumann K
Characterization of two distinct subfamilies of SUN-domain proteins in Arabidopsis and their interactions with the novel KASH-domain protein AtTIK.
Published on 30 Dec 2014 in Journal of experimental botany , vol. 65 - pp 6499-512
Graumann K, Vanrobays E , Tutois S , Probst AV , Evans DE, Tatout C
Evolutionary history of Methyltransferase 1 genes in hexaploid wheat.
Published on 23 Oct 2014 in BMC genomics , vol. 15 - pp 922
Thomas M, Pingault L, Poulet A , Duarte J, Throude M, Faure S, Pichon JP, Paux E, Probst AV , Tatout C
The plant LINC complex at the nuclear envelope
Published on 30 Jun 2014 in Chromosome research , vol. 22 - pp 241-52
Tatout C , Evans DE, Vanrobays E , Probst AV , Graumann K
Heterochromatin reorganization during early mouse development requires a single-stranded noncoding transcript.
Published on 26 Sep 2013 in Cell reports , vol. 4 - pp 1156-67
Casanova M, Pasternak M, El Marjou F, Le Baccon P, Probst AV , Almouzni G
Heterochromatin dynamics during developmental transitions in Arabidopsis – a focus on ribosomal DNA loci.
Published on 15 Aug 2013 in Gene , vol. 526 - pp 39-45
Benoit M , Layat E , Tourmente S , Probst AV
Transcript levels, alternative splicing and proteolytic cleavage of TFIIIA control 5S rRNA accumulation during Arabidopsis thaliana development.
Published on 30 Jul 2012 in The Plant journal : for cell and molecular biology , vol. 71 - pp 35-44
Layat E , Cotterell S , Vaillant I , Yukawa Y, Tutois S , Tourmente S
The N-terminal domains of TRF1 and TRF2 regulate their ability to condense telomeric DNA.
Published on 30 Mar 2012 in Nucleic acids research , vol. 40 - pp 2566-76
Poulet A, Pisano S, Faivre-Moskalenko C, Pei B, Tauran Y, Haftek-Terreau Z, Brunet F, Le Bihan YV, Ledu MH, Montel F, Hugo N, Amiard S , Argoul F, Chaboud A, Gilson E, Giraud-Panis MJ
Heterochromatin maintenance and establishment: lessons from the mouse pericentromere.
Published on 01 Sep 2011 in Nucleus (Austin, Tex.) , vol. 2 - pp 332-8
Almouzni G, Probst AV
Characterization of a cinnamoyl-CoA reductase 1 (CCR1) mutant in maize: effects on lignification, fibre development, and global gene expression.
Published on 30 Jul 2011 in Journal of experimental botany , vol. 62 - pp 3837-48
Tamasloukht B, Wong Quai Lam MS, Martinez Y, Tozo K, Barbier O, Jourda C, Jauneau A, Borderies G, Balzergue S, Renou JP, Huguet S, Martinant JP, Tatout C , Lapierre C, Barrière Y, Goffner D, Pichon M
Heterochromatin establishment in the context of genome-wide epigenetic reprogramming.
Published on 30 May 2011 in Trends in genetics : TIG , vol. 27 - pp 177-85
Probst AV , Almouzni G
SUMOylation promotes de novo targeting of HP1α to pericentric heterochromatin.
Published on 30 Mar 2011 in Nature genetics , vol. 43 - pp 220-7
Maison C, Bailly D, Roche D, Montes de Oca R, Probst AV , Vassias I, Dingli F, Lombard B, Loew D, Quivy JP, Almouzni G
A strand-specific burst in transcription of pericentric satellites is required for chromocenter formation and early mouse development.
Published on 19 Oct 2010 in Developmental cell , vol. 19 - pp 625-38
Probst AV , Okamoto I, Casanova M, El Marjou F, Le Baccon P, Almouzni G
TRF2 and apollo cooperate with topoisomerase 2alpha to protect human telomeres from replicative damage.
Published on 23 Jul 2010 in Cell , vol. 142 - pp 230-42
Ye J, Lenain C, Bauwens S, Rizzo A, Saint-Léger A, Poulet A, Benarroch D, Magdinier F, Morere J, Amiard S , Verhoeyen E, Britton S, Calsou P, Salles B, Bizard A, Nadal M, Salvati E, Sabatier L, Wu Y, Biroccio A, Londoño-Vallejo A, Giraud-Panis MJ, Gilson E
Platination of telomeric DNA by cisplatin disrupts recognition by TRF2 and TRF1.
Published on 30 Jun 2010 in Journal of biological inorganic chemistry : JBIC : a publication of the Society of Biological Inorganic Chemistry , vol. 15 - pp 641-54
Ourliac-Garnier I, Poulet A, Charif R, Amiard S , Magdinier F, Rezaï K, Gilson E, Giraud-Panis MJ, Bombard S
Identification and mapping of induced chromosomal deletions using sequence polymorphisms.
Published on 30 Jan 2010 in BioTechniques , vol. 48 - pp 53-60
Vanrobays E , Jennings BH, Ish-Horowicz D
Heterochromatin at mouse pericentromeres: a model for de novo heterochromatin formation and duplication during replication.
Published on 01 Jan 2010 in Cold Spring Harbor symposia on quantitative biology , vol. 75 - pp 155-65
Maison C, Quivy JP, Probst AV , Almouzni G
A Pol V-mediated silencing, independent of RNA-directed DNA methylation, applies to 5S rDNA.
Published on 30 Oct 2009 in PLoS genetics , vol. 5 - pp e1000690
Douet J, Tutois S , Tourmente S
TRF2 promotes, remodels and protects telomeric Holliday junctions.
Published on 18 Mar 2009 in The EMBO journal , vol. 28 - pp 641-51
Poulet A, Buisson R, Faivre-Moskalenko C, Koelblen M, Amiard S , Montel F, Cuesta-Lopez S, Bornet O, Guerlesquin F, Godet T, Moukhtar J, Argoul F, Déclais AC, Lilley DM, Ip SC, West SC, Gilson E, Giraud-Panis MJ
TOR regulates the subcellular distribution of DIM2, a KH domain protein required for cotranscriptional ribosome assembly and pre-40S ribosome export.
Published on 30 Oct 2008 in RNA (New York, N.Y.) , vol. 14 - pp 2061-73
Vanrobays E , Leplus A, Osheim YN, Beyer AL, Wacheul L, Lafontaine DL
Hypomethylation and hypermethylation of the tandem repetitive 5S rRNA genes in Arabidopsis.
Published on 30 Apr 2008 in The Plant journal : for cell and molecular biology , vol. 54 - pp 299-309
Vaillant I , Tutois S , Jasencakova Z, Douet J, Schubert I, Tourmente S
In Drosophila melanogaster the COM locus directs the somatic silencing of two retrotransposons through both Piwi-dependent and -independent pathways.
Published on 06 Feb 2008 in PloS one , vol. 3 - pp e1526
Pericentric heterochromatin: dynamic organization during early development in mammals.
Published on 30 Jan 2008 in Differentiation; research in biological diversity , vol. 76 - pp 15-23
Probst AV , Almouzni G